How Do We Measure Antimicrobial Resistance?
ID
HORT-258NP
These days antimicrobial resistance (AMR) is prominent in the headlines, with growing concern that these life-saving drugs will continue to lose effectiveness. While there is growing research effort working towards understanding and stopping the spread of AMR, the practical application of such research can be difficult to grasp. A key issue to interpreting research reports about AMR is understanding the different ways that it is measured. This guide will help you better understand what scientists specifically mean when they report their findings about AMR.
What is the difference between an antibiotic, ARB, and ARG?

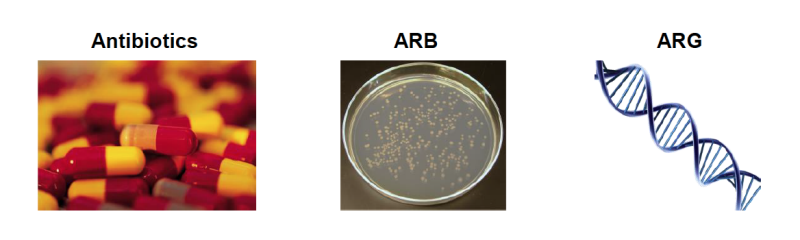
Antibiotics: Antibiotics are drugs given to livestock or humans, such as penicillin or erythromycin, in order to treat or prevent a bacterial infection. New scientific methods make it possible to detect residual antibiotics in manure, sewage, and affected water, soil, and other environments. Different antibiotics are used to treat and prevent different kinds of bacterial infections.
Antibiotic Resistant Bacteria (ARB): Antibiotic resistant bacteria are able to survive and even grow in the presence of antibiotics. ARBs can be “good” (e.g., normal flora) or “bad” (e.g., pathogens) bacteria. While pathogenic ARBs are of most concern, non-pathogens are also important because they can pass along the ability to resist antibiotics to other bacteria.
Antibiotic Resistance Gene (ARG): An ARG is a part of an ARB’s DNA that encodes the ability for it to live in the presence of antibacterial drugs. ARGs are of concern because they can transfer between bacteria. In other words, bacteria that are not resistant can receive an ARG from another bacteria and become resistant.
Scientific reports typically provide data about how much of an antibiotic, ARB, or antibiotic resistance gene (ARG) was measured in a target sample. The implication is that higher levels of antibiotics, ARBs, or ARGs indicate a higher level of risk of spreading AMR and more individuals contracting resistant infections. However, a current scientific challenge is that we do not know how many antibiotics, ARBs, or ARGs is a real risk.
As of 2017, there are no published guidelines that tell us what amount of antibiotics, ARBs, or ARGs equal a human health risk.
How do Scientists Measure AMR?
Researchers can use a variety of techniques to determine if antibiotics or ARBs are present. They can grow bacteria in the lab or use state-of-the-art technology to look at DNA.
In the lab, researchers can spread a sample on a culture dish containing nutrients formulated specifically to help the bacteria grow. To figure out which bacteria are resistant, antibiotics can be added to the culture media or by placing small discs infused with different antibiotics on the media surface (i.e., Kirby Bauer method). Plates are examined to see if bacterial colonies were able to grow in the presence of the antibiotics. ARBs grow on media containing the antibiotic or in close proximity to the discs.
The bacterial DNA can also be studied in order to search directly for ARGs. Researchers can directly isolate bacterial DNA from manure, soil, water, and other environmental samples. They can then compare the DNA to databases used by researchers all around the world in order to determine what kinds of ARGs may be present. This method is known as metagenomics. As of now, there are hundreds to thousands of known ARGs, with attention to any new ARGs that could enable pathogens to be resistant of special interest.
These are just a couple of the many ways antimicrobial resistance is studied by scientists. Understanding these concepts can help better interpret data presented by the media and in scientific reports and how the findings have real-world application.
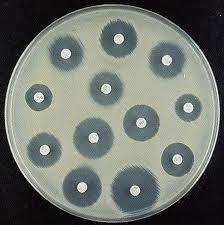
Kirby Bauer Test
After spreading the sample, discs containing different antibiotics are gently placed on the surface of the media. The clearings around the discs are where bacteria in the sample could not grow because of the antibiotic.
References
Information on antibiotic resistant bacteria (ARB) and antibiotic resistance genes (ARGs). Available from: http://emerald.tufts.edu/med/apua/about_issue/about_antibioticres.shtml. Accessed July 14, 2017.
Kirby Bauer Test-Information. Available from: https://www.boundless.com/microbiology/textbooks/boundless-microbiology-textbook/antimicrobial-drugs-13/measuring-drug-susceptibility-157/kirby-bauer-disk-susceptibility-test-791-6152/. Accessed July 14, 2017.
What is Metagenomics? Available from: http://www.news-medical.net/life-sciences/What-is-Metagenomics.aspx. Accessed July 14, 2017.
Virginia Cooperative Extension materials are available for public use, reprint, or citation without further permission, provided the use includes credit to the author and to Virginia Cooperative Extension, Virginia Tech, and Virginia State University.
Virginia Cooperative Extension is a partnership of Virginia Tech, Virginia State University, the U.S. Department of Agriculture (USDA), and local governments, and is an equal opportunity employer. For the full non-discrimination statement, please visit ext.vt.edu/accessibility.
Publication Date
March 29, 2023



